Achieve Unparalleled Structural Variant Detection
Routinely detect all classes of structural variants down to 500 base pair (bp) resolution and 5% variant allele frequency (VAF) with optical genome mapping (OGM) using the Saphyr™ system. Saphyr™ is the most powerful structural variant detection tool available, detecting genomic variants commonly missed by next-generation sequencing (NGS) technologies and conventional cytogenetic techniques.


Saphyr™ System Brochure
Explore the Saphyr system brochure to get an overview of the system, range of applications, workflow and more.
DownloadEmpower your lab with advanced SV detection using optical genome mapping.
The Saphyr™ system provides optical genome mapping of ultra-high molecular weight (UHMW) DNA in its native state, from molecules 150 kbp to multi-megabase pairs in length.
In as little as three days, complete the entire workflow from sample to structural variation call using the Bionano sample preparation kits, Saphyr™ system, and software.

Achieve broad, unbiased genomic coverage with flexible data collection.
The genomic coverage achieved with the Saphyr™ system is flexible and allows for the detection of heterozygous variants or rarer variants found in mosaic samples and heterogeneous tumors.
Collect 100X coverage of a human genome in as little as six hours. See deeper and rarer variants by simply extending the data collection time on Saphyr™ at no additional consumables cost. Achieve 400X coverage in 24 hours and provide SV detection down to 5% VAF. Further extend runs to see even lower VAF.
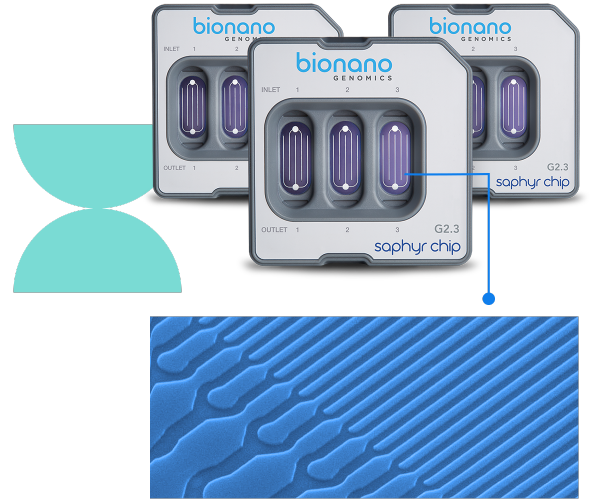
Experience seamless integration and workflow using the Saphyr™ Chip® consumable and Saphyr™ system.
The Saphyr Chips utilize hundreds of thousands of massively parallel nanochannels that linearize long, labeled DNA molecules, allowing the Saphyr instrument to directly image your samples.
Sample loading on the Saphyr Chip is fast and easy. Pipet up to three samples into individual flow cells on the chip. Load up to two chips onto the Saphyr instrument per run for a maximum throughput of six samples per run.
The Saphyr instrument and chips leverage machine learning-based adaptive loading and automatic run optimization to ensure walk-away scanning and the fastest possible throughput for runs.

Ensure maximum performance with automated health system monitoring via remote access.
The Saphyr™ Assure Service provides an optional (opt-in) automated health monitoring feature that continuously inspects data quality and instrument performance. The feature detects issues before they effect data quality and performance so Bionano Support can proactively repair the system with minimal downtime.
Saphyr™ System Performance by Application
The performance details of Saphyr™ vary depending on the application and analysis pipeline used.
| Application | Germline DNA Analysis | Cancer Analysis |
| Data collected | 400 Gbp | 1.5 Tbp |
| Coverage setting | 100x | 400x |
| Effective coverage | 80x | 300x |
| Variant allele frequency | 50% | >5% |
| Analysis pipeline | De novo assembly | Rare variant analysis |
| Performance (size) by variant type* | ||
| Insertions | >500 bp | >5 kbp |
| Deletions | >700 bp | >7 kbp |
| Stable Repeat Expansion / Contractions | >500 bp | >5 kbp |
| Duplications | >30 kbp | >150 kbp |
| Translocations | >70 kbp | >70 kbp |
| Inversions | >30 kbp | >70 kbp |
*Performance shown is >90% sensitivity
Learn more about OGM
Read about what structural variations are and why they matter.
Learn MoreSee how OGM reveals structural variation in a way that has never been done before.
Learn MoreFind the latest research in our Publications Library.
Learn MoreHear What Some of our Customers Say About Using the Saphyr™ OGM System in Their Research
Downloads and Resources
Explore all the benefits of the Saphyr™ system
Review product ordering information
See detailed information about how Saphyr™ works

Speak to an expert, to learn more details on how OGM works and how it can fit the needs of your institution
Talk to a specialistRelated Products
