Blog
Significant Cost Savings with Single NGS Test vs. Other Strategies
A few months ago, we shared a conversation with Dr. Ingrid Simonic, Medical Director at Cambridge University Hospitals, who talked about the benefits of consolidating multiple testing strategies into a single NGS assay. She mentioned that the large number of tests performed are not directly correlated to the number of patients but that multiple tests are performed per patient in sequence. She then took the challenge to see how they could consolidate their testing practices to save time and improve yield and concluded that a single CNV/SNV pipeline (vs. multi-tiered tests) would streamline the process providing faster results.
In that post, the bottom line was that a single CNV/NGS assay is beneficial over multiple tests, increasing diagnostic yield while saving time. Here we examine how NGS testing (a single test) as opposed to sequential gene testing (multiple tests) results in significant cost savings. At the American Society of Clinical Oncology (ASCO) meeting last month, Dr. Nathan Pennell, Director of the Lung Cancer Medical Oncology Program at the Cleveland Clinic, and his colleagues presented cost analysis in a model with different types of genetics testing in metastatic non-small cell lung cancer (mNSCLC). The team showed that upfront NGS leads to significant cost savings vs. sequential single-gene testing modalities.
NSCLC Testing
There are many tests offered for NSCLC but no single standard followed by all. Current NCCN guidelines for treatment planning for NSCLC advise looking at PD-L1 expression and mutations in genes linked to NSCLC. PD-L1 (programmed cell death ligand) prevents T cells from attacking cancer cells. There are a number of approved drugs that block PD-L1 from attaching to T cells thereby allowing the cells to perform their natural task of attacking cancer cells. EGFR, ALK, ROS1, BRAF, MET, HER2, RET, NTRK1 are genes known to have mutations in NSCLC patients. Of these, EGFR, ALK, ROS1, BRAF have approved therapies while the other genes have investigational or off-label agents for treatment.
Study Design
The study used a model of a million members in an insurance plan and estimated 2,066 Center for Medicare and Medicaid Services (CMS) and 156 commercially insured mNSCLC patients would receive PD-L1 and genomic alterations (GA) tests using one of four strategies:
- Sequential tests (one GA test at a time; one gene at a time)
- Exclusionary mutation (KRAS) test followed by sequential tests (if KRAS is negative, sequential tests would follow); KRAS is the most common mutation so if this were positive, these patients could be excluded from further testing
- Panel test – all single GA tests performed at the same time
- Upfront NGS – a single test including all genes from the GA tests plus KRAS
Of the known biomarkers linked to NSCLC, the study first considered the genes with approved therapies and if nothing was found there, then looked at the other genes. Groups 1-3 above were tested for mutations in genes with approved treatment followed by single gene testing or panel for the other genes. The amount of time and the cost for testing was evaluated. The group also accounted for the need for additional biopsies into their calculations.
Study Results
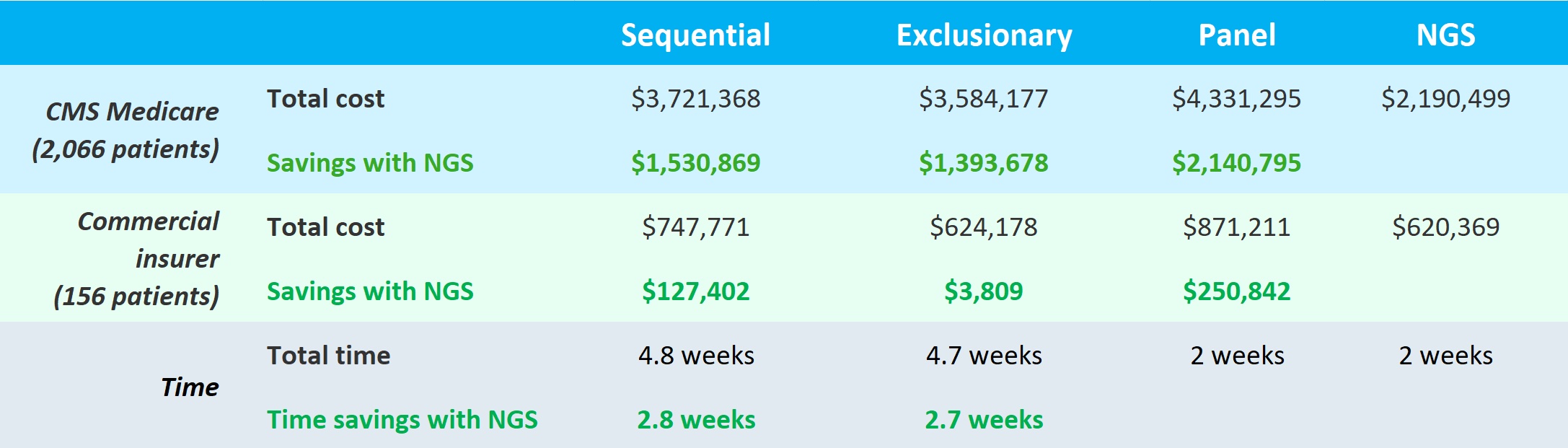
Panel and NGS testing took the same amount of time but were faster by 2.7 and 2.8 weeks than exclusionary and sequential, respectively. But NGS came out the clear winner for both CMS and commercially insured in terms of costs. Use of NGS saved roughly 1.4 to 2.1 million dollars for CMS and for commercial insurers, ~$3,800 to ~$250,000 over the other tests. Dr. Pinnell stated that they were able to identify more patients with actionable mutations with the single upfront NGS test than other methods. So in the end, the single NGS test was better for both patients (faster and higher diagnostic yield) and payors (significant cost savings).
This was an economic model study and the group would like to do additional studies using real-world data. Also, this study examined only sequence variants using NGS in a specific disease. But one can expect even greater savings when the NGS test is also used to derive CNVs in addition to sequence variants as promoted by Dr. Simonic.
